|
Hello! I am a second-year Ph.D. student in the Neural Computation & Machine Learning joint Ph.D. program at Carnegie Mellon University supported by an NSF Graduate Research Fellowship. I am advised by Prof. Michael Tarr and Prof. Deva Ramanan. My primary interests lie in understanding how humans understand dynamic visual stimuli and improving machine visual reasoning. I graduated with a B.S. in Electrical Engineering and Computer Science (EECS) from UC Berkeley. My research was advised by Dr. Kristofer Bouchard and Dr. Ji Hyun Bak at the Lawrence Berkeley National Laboratory. I also worked with Medhini Narasimhan and Prof. Trevor Darrell at Berkeley AI Research (BAIR). |
|
|
|

|
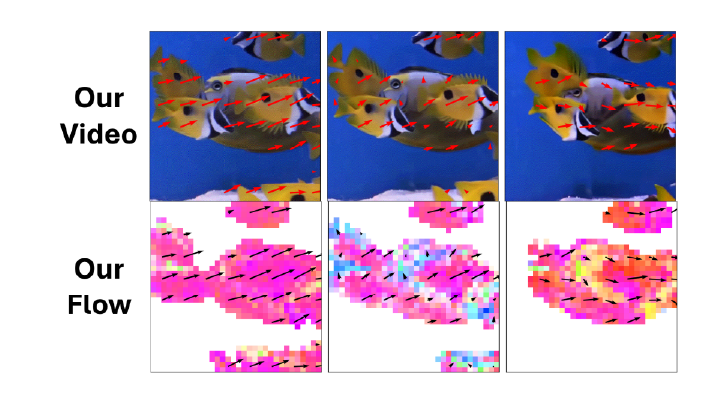
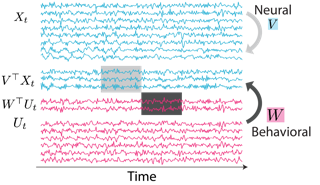
Jacob Yeung, Andrew F. Luo, Gabriel Sarch, Margaret M. Henderson, Deva Ramanan, Michael J. Tarr CVPR 2025 Oral, (top 0.7% of all submissions) Paper / Website / bibtex We propose a way to study visual motion in the brain by decoupling the modeling of static image representations and motion representations in the human brain. |
|
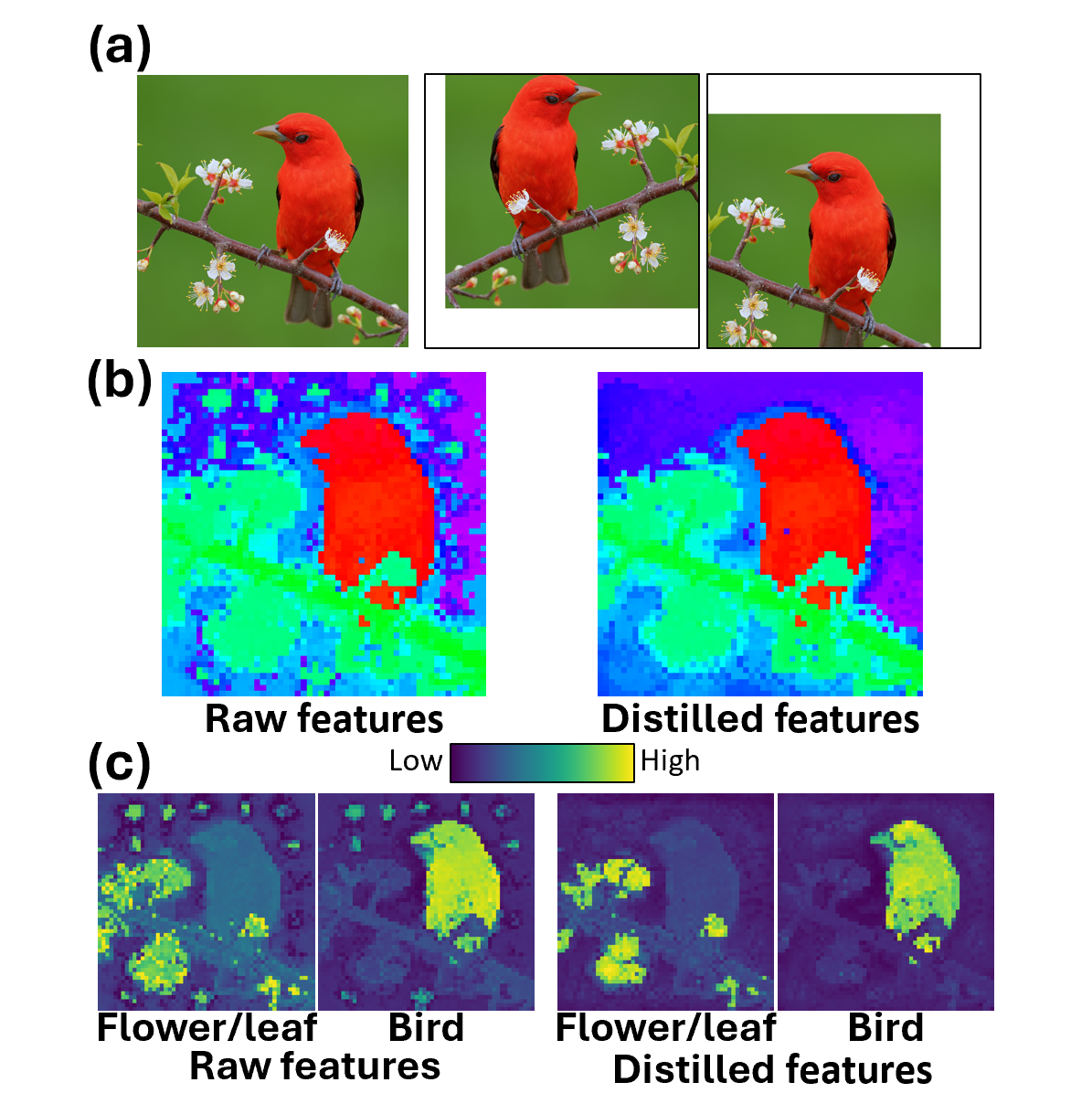
Andrew F. Luo, Jacob Yeung, Rushikesh Zawar, Shaurya Dewan, Margaret M. Henderson, Leila Wehbe, Michael J. Tarr ICLR 2025 Paper / bibtex We propose an efficient gradient-free distillation module capable of extraction high quality dense CLIP embeddings, and utilize these embeddings to understand semantic selectivity in the visual cortex. |
|
Jacob Yeung, Ji Hyun Bak, Kristofer Bouchard COSYNE, 2023 Paper / Poster We introduce external Dynamics Components Analysis (eDCA), a linear dimensionality reduction method that finds subspaces of past neural data that have maximal mutual information with subspaces of future behavior data. |
|
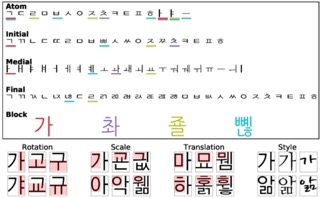
Jesse Livezey, Ahyeon Hwang, Jacob Yeung, Kristofer Bouchard International Conference on Image Analysis and Processing, 2022 Paper / bibtex We introduce a dataset with known hierarchical and compositional structure and show deep unsupervised models poorly learn the latent generative structures. |
|
|
|
I have been a teaching assistant for the following courses: |
|
I have been a tutor for the following courses: |
|
|